
CASPER (CRISPR Associated Software for Pathway Engineering and Research) is a flexible GUI-based software platform for gRNA generation and analysis in any organism and with any CRISPR-Cas system. Based on the algorithms described in a recent Trinh Lab publication, CASPER combines traditional gRNA design tools with unique functions like:
- Multiple-Cas-type gRNA generation
- Spacer redundancy evaluation in a single species or microbiome
- Programmable on- and off-target score weighting
CASPER is available on Github and may be used on all major operating systems.
Tutorials for CASPER can be found on the Trinh Lab YouTube channel.
If you have any questions about CASPER, please open an issue on Github or email David Dooley.
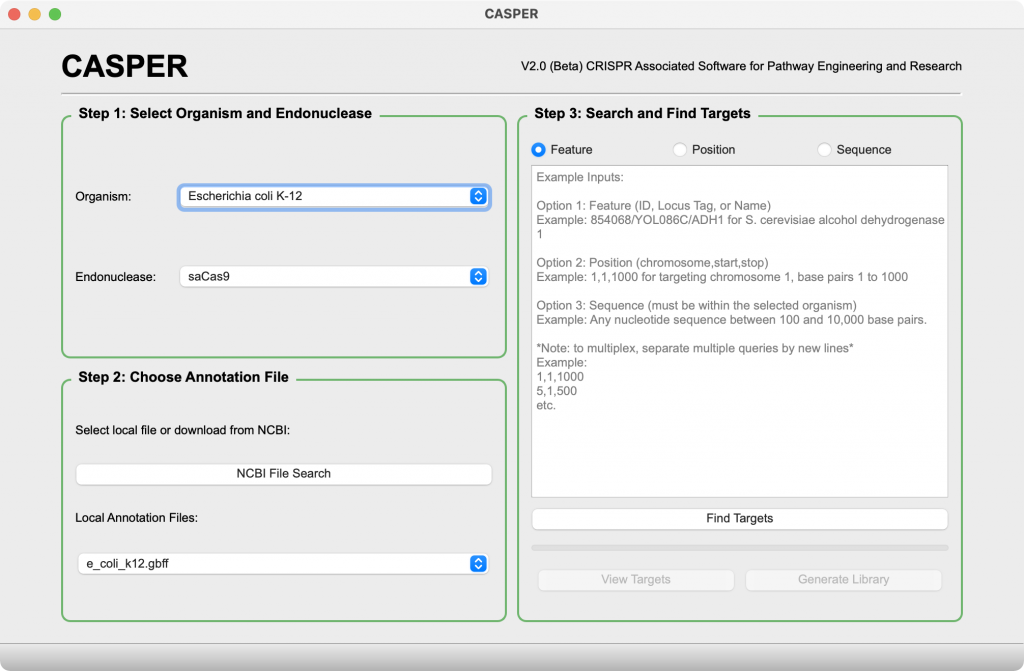
Main Window
- When paired with a GenBank formatted annotation file, users can search for gRNA targets using gene name, ID, locus tag, position, and even sequence.
- Seamless integration with NCBI’s public databases allows users to query and download genomic sequence and annotation files directly from CASPER.
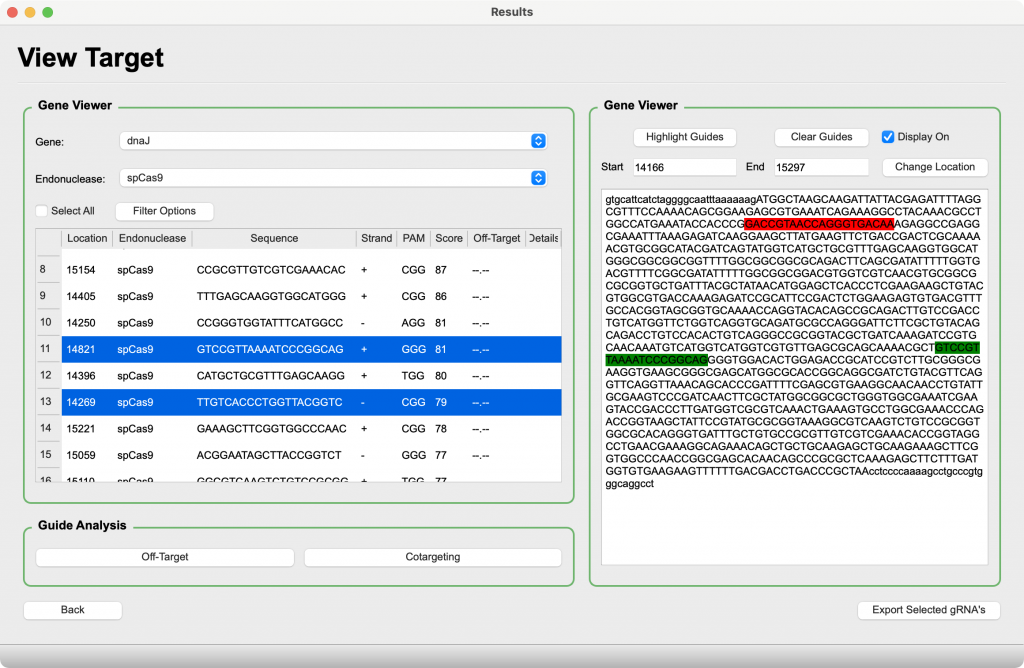
View Target
CASPER allows users to filter gRNAs to find targets that really work!
- On- and off-target scoring
- 5’ G filtering for expression from T7 promoter
- Minimum on-target score
- Cotargets only
Gene Viewer function allows users to see exactly where their gRNAs localize within their gene of interest.
Multitargeting
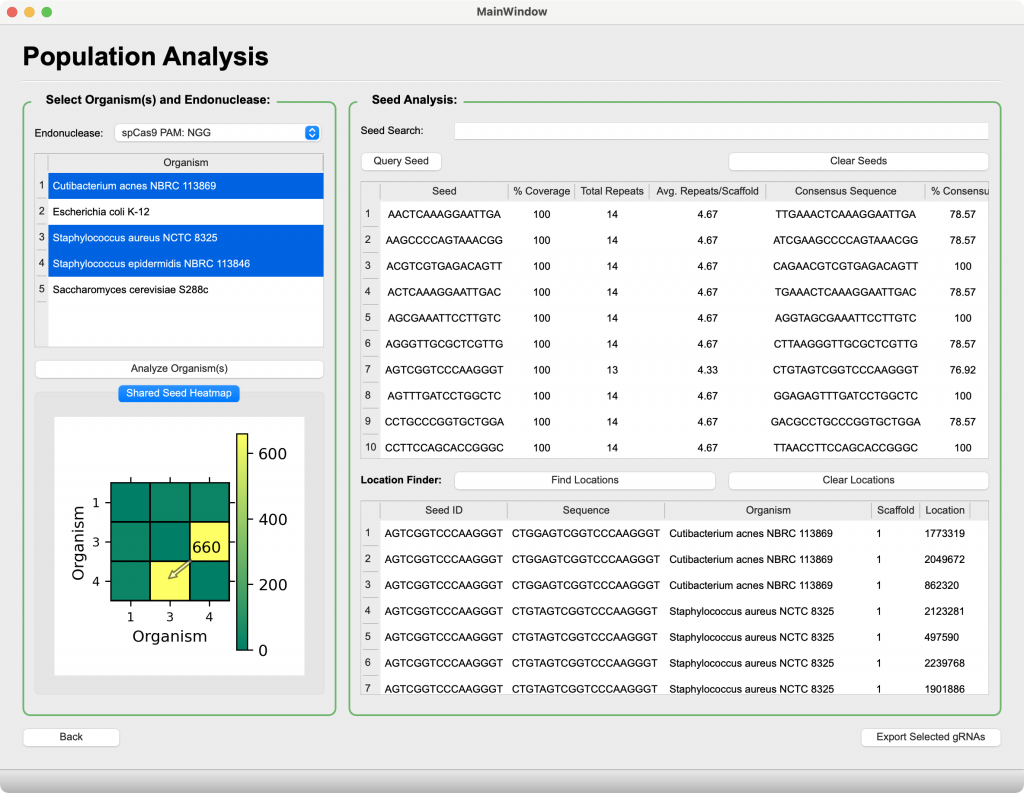
Population Analysis
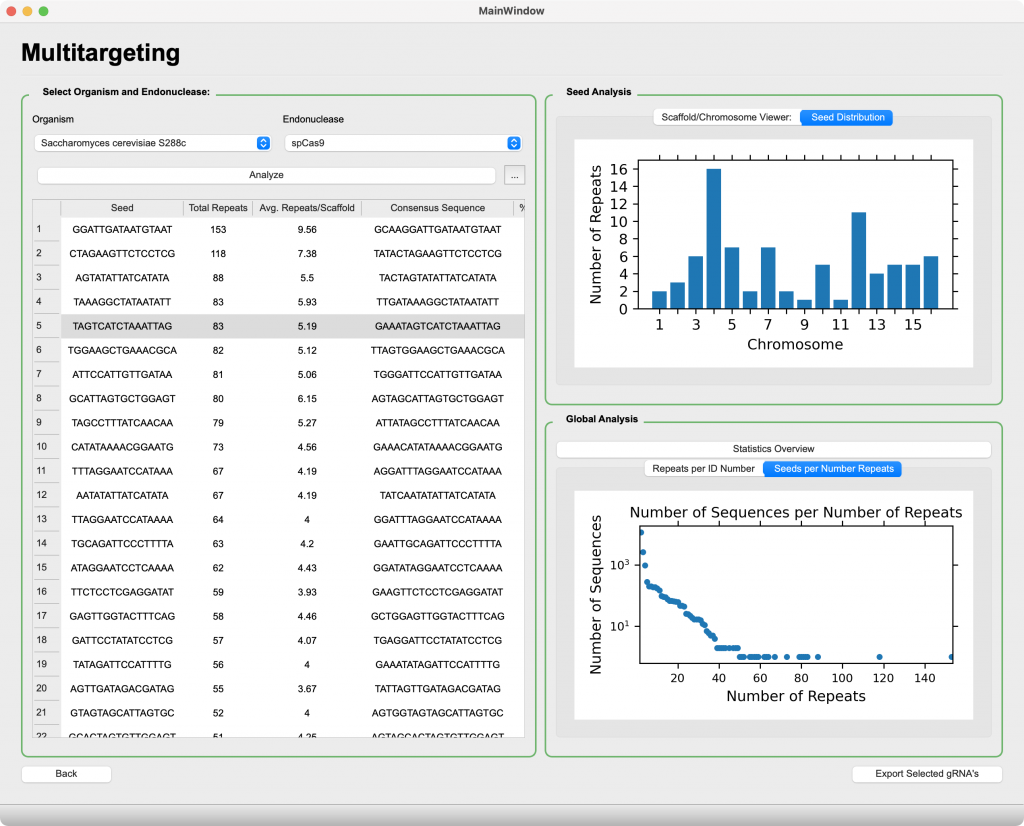
Maximize efficiency of your CRISPR assays by designing single gRNAs active at multiple sites
- Analyze and visualize redundant gRNA motifs across entire genomes
- Global statistics enable high-level characterization of an organism’s multitargeting potential
Enabling tool for CRISPR-based microbiome manipulation
- View pairwise conserved gRNA sequences between several organisms within a metagenome
- Determine globally conserved targets across an entire population
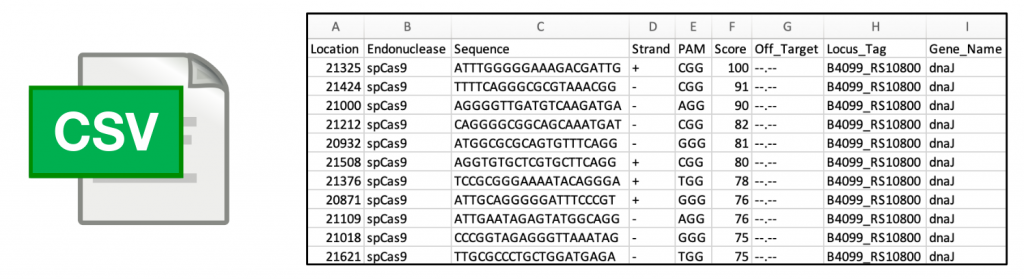
Export to CSV
At every step, gRNAs can be exported to a CSV file, making further manipulation and ordering of your targets trivial with your favorite sheet-based editor.
